by David Rawlinson and Gideon Kowadlo
A Temporal Pooler Proposal
We will describe a Temporal Pooler design below. We have tested this design and it gives satisfactory output given the criteria and expectations described below and in the previous articles in this series (post #1, post #2). It is also relatively simple. You could imagine this design being implemented in biology, but we have no biological evidence that this is the design used in the cortex. Nevertheless, it is useful to have a template for an "ideal" design (from a certain perspective), against which to compare known biology.
[EDIT: We have now rejected this design and prefer temporal pooling via on Predictive Coding]
This article will first list our design criteria, then give pseudocode for the TP implementation. Finally we will work through some examples to show the output of our temporal pooler.
Assumptions
1. This is a design based on engineering requirements, not based on biology
2. For simplicity of explanation, we will assume a "Region" is a piece of cortex comprising several layers of cells with distinct functions. There is mutual inhibition and high internal connectivity within the region. The region has a finite 2-d extent within all layers.
3. For a given set of active feed-forward (FF) input cells (let's call this an input "pattern"), each sub-region will have a single "winning" spatial-pooler (SP) cell that most closely matches the input pattern. This differs from the CLA/HTM definition of a region, where there may be multiple "winners". However, for this example let's keep things simple.
4. The finite set of cells local to the region mean that all configurations of world and agent are represented by states in a small, loopy graph embodied by the region's SP cells. "Loopy" means that there is at least one cycle and that there are no terminal states in this digraph. We understand that the local representation of agent + world state will be missing a lot of detail, that that's OK.
5. The temporal pooler will only update on a change to the FF input pattern.
6. Temporal Pooling should guarantee to increase output stability for any given sequence of SP cell activation. This means that sequences of length at least 3 must be replaced with constant output from the Temporal Pooler.
7. Uncertainty should not be hidden by the Temporal Pooler. Where forks occur in the graph, TP output should change, unless the fork is predictable. If an unpredicted event occurs, TP output should change to allow the transition to be modelled at a higher hierarchy level.
8. We assume that state-splitting creates redundant SP-cells that respond to the same FF input, but with different prior (previous) cell activations. e.g. SP cell #1 responds to uniquely to "green" after "blue", and SP cell #2 responds to "green" after "red". See the earlier post for more on this. A combination of state-splitting and simultaneous multiple TP-cell activation is necessary to guarantee pooling. It also allows variable-order modelling of FF input in different sequential contexts.
Outline
We require 2 sets of cells for our design. First, a layer of SP cells. A single SP cell fires for a given FF input, and represents the observation of that input when active.
Second, a layer of TP cells. We require 1 TP cell per SP cell. While SP cells fire individually, and only one at any given time, a set of TP cells will be simultaneously active. The set of active TP cells collectively represent a sequence of FF input, and the set must collectively fire only on observation of a specific and unique sequence of active SP cells. In a hierarchy, the next layer's SP cells will replace the active TP cells with individual cells that represent these combinations.
At all times at least 1 TP cell will be active.
The design has 2 key elements:
- Propagation of TP cell activity based on first-order modelling of historical transitions between SP cells.
- Assumes state splitting has created redundant SP cells representing the same FF input patterns, thereby allowing variable-order modelling using only first-order relations between SP cells.
- Inhibition of TP cell activity after a prediction failure.
Pseudocode
Variables:
nCells: The number of SP and TP cells
spCells: An array of SP cell activation values, length nCells
inCells: An array of TP cell inhibition values, length nCells
tpCells: An array of TP cell activation values, length nCells
spBest: The winning SP cell that matches the current FF input
spPrev: The previous winning SP cell.
spPred: The SP cell predicted to be the next best cell.
w( i, j ): Probability of transition from SP cell i to SP cell j in local graph
Note that weights w(i,j) represent conditional probabilities w(i,j) = P( S'=j | S=i ) where S is the currently active SP cell and S' is the next active SP cell. The weights can be approximated using historical observations of transition frequencies between active SP cells. Learning of sequence memory transitions is not covered in the pseudocode below. Similarly, the process for learning and defining the set of SP cells is not described.
// clear tp cell activity after bad prediction
if( spPred != spBest ) { // last prediction wrong
inCells[ spPrev ] = 1;
tpCells[ all ] = 0;
}
// clear tp cell activity when we reach inhibited cell
if( inCells( spBest ) ) {
tpCells[ all ] = 0;
}
// clear inhibition of winning cell and activate it
inCells[ spBest ] = 0
tpCells[ spBest ] = 1
// now propagate this activity forwards
Figure 3: Say that via state-splitting, we have constructed a set of 7 SP cells (centre graph, coloured and numbered circles). This means we also need to create 7 TP cells (we specified earlier that there would be 1 TP cell per SP cell). Each TP cell is associated with one SP cell. The magenta outline of SP cell 1 indicates that this cell has been selected as best matching the current FF input. None of the TP cells are active. The heavier arrow from SP cell 1 to 2 represents the higher probability of this outcome.
Figure 4: According to the algorithm given above, we will first activate TP cell 1 due to activity of SP cell 1 (magenta). Then, we propagate forwards the activation through TP cells 2,3,4 because this is the most likely path (brown outline highlights). Activation propagates back to TP cell 1 and then has no further effect, since TP cell 1 is already active. TP cells 5,6,7 are not activated because this is not the most likely path from TP cell 1. As a result, of our 7 TP cells, 4 are active {1,2,3,4} of {1,2,3,4,5,6,7}.
Figure 5: The external world moves from a state generating the black observation, to green. SP cell 2 is correspondingly made active. The temporal pooler cells are now updated, starting by activating TP cell 2 (although it is already active). TP cell 1 remains active for two reasons: First, activity is not cleared unless there is a prediction failure. Second, activity propagates from TP cell 2 through TP cells 3,4, and 1, back to 2. Again, TP cells 5,6,7 are not active. There has been no change in the set of active TP cells as they are following a predictable sequence. In fact, the set of active TP cells will not change while visiting any of the active states {1,2,3,4}.
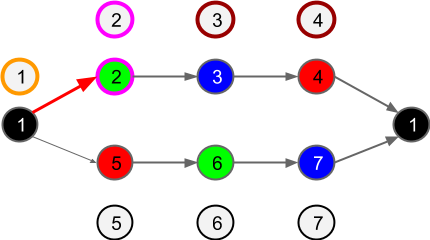
Figure 6: If, randomly, a transition from SP cell 1 to SP cell 5 occurs, the pattern of TP cell activation will change. TP cell 1 will be inhibited (magenta X) due to the prediction failure. TP cell 5 will be activated and the activity will propagate through TP cells 6 and 7. Crucially, it cannot propagate further due to inhibition of TP cell 1. This prevents the set of active cells extending to TP cells 2,3,4.
Figure 7: For the remainder of the sequence R-G-B, TP cells 5,6,7 remain active. While Green is observed, as shown above, TP cell 5 remains active not due to forward prediction but due to the fact that TP cell activity is retained until there is a prediction failure.
Discussion
In this example, TP output is one of two subsets of cells, either {1,2,3,4} or {5,6,7}. In the next hierarchy layer, the active subsets would be associated with specific SP cells that would represent these sequences of observations in their entirety.
This is a relatively simple scheme that only requires inhibition and prediction. A prediction failure is always followed by a change in TP output, due to the inhibition mechanism, even if the most likely path leads back to the original set of active cells prior to the failed prediction. Recall that the local graph being modelled in a region of cortex will always feature cycles and have no terminal states. Without inhibition, we would end up with scenarios where all TP cells were constantly active regardless of which fork was taken (if a lower-probability outcome occurs). This would obscure the unpredictable transitions and make them unavailable for modelling higher in the hierarchy.
At least 1 TP cell is always active (the cell that is associated with the currently active SP cell).
The output of the temporal pooler is maximally stable when sequences of observations are predictable. This is ideal if we assume the predicted outcome is the most common outcome. The propagation mechanism as proposed makes best use of predictable transitions - sequences of arbitrary length can be replaced with a constant output, if the system is sufficiently predictable.
Note that this method can only guarantee to simplify the observed problem in combination with state-splitting. The latter is required to build more and longer deterministic sequences of SP cell activation, even if these sequences are not predictable. It is assumed that given increasingly higher-order interpretation, predictability will be achieved.
In a future post we will look in more detail about the process of state-splitting.
A Temporal Pooler Proposal
We will describe a Temporal Pooler design below. We have tested this design and it gives satisfactory output given the criteria and expectations described below and in the previous articles in this series (post #1, post #2). It is also relatively simple. You could imagine this design being implemented in biology, but we have no biological evidence that this is the design used in the cortex. Nevertheless, it is useful to have a template for an "ideal" design (from a certain perspective), against which to compare known biology.
[EDIT: We have now rejected this design and prefer temporal pooling via on Predictive Coding]
This article will first list our design criteria, then give pseudocode for the TP implementation. Finally we will work through some examples to show the output of our temporal pooler.
Assumptions
1. This is a design based on engineering requirements, not based on biology
2. For simplicity of explanation, we will assume a "Region" is a piece of cortex comprising several layers of cells with distinct functions. There is mutual inhibition and high internal connectivity within the region. The region has a finite 2-d extent within all layers.
3. For a given set of active feed-forward (FF) input cells (let's call this an input "pattern"), each sub-region will have a single "winning" spatial-pooler (SP) cell that most closely matches the input pattern. This differs from the CLA/HTM definition of a region, where there may be multiple "winners". However, for this example let's keep things simple.
4. The finite set of cells local to the region mean that all configurations of world and agent are represented by states in a small, loopy graph embodied by the region's SP cells. "Loopy" means that there is at least one cycle and that there are no terminal states in this digraph. We understand that the local representation of agent + world state will be missing a lot of detail, that that's OK.
5. The temporal pooler will only update on a change to the FF input pattern.
6. Temporal Pooling should guarantee to increase output stability for any given sequence of SP cell activation. This means that sequences of length at least 3 must be replaced with constant output from the Temporal Pooler.
7. Uncertainty should not be hidden by the Temporal Pooler. Where forks occur in the graph, TP output should change, unless the fork is predictable. If an unpredicted event occurs, TP output should change to allow the transition to be modelled at a higher hierarchy level.
8. We assume that state-splitting creates redundant SP-cells that respond to the same FF input, but with different prior (previous) cell activations. e.g. SP cell #1 responds to uniquely to "green" after "blue", and SP cell #2 responds to "green" after "red". See the earlier post for more on this. A combination of state-splitting and simultaneous multiple TP-cell activation is necessary to guarantee pooling. It also allows variable-order modelling of FF input in different sequential contexts.
Outline
We require 2 sets of cells for our design. First, a layer of SP cells. A single SP cell fires for a given FF input, and represents the observation of that input when active.
Second, a layer of TP cells. We require 1 TP cell per SP cell. While SP cells fire individually, and only one at any given time, a set of TP cells will be simultaneously active. The set of active TP cells collectively represent a sequence of FF input, and the set must collectively fire only on observation of a specific and unique sequence of active SP cells. In a hierarchy, the next layer's SP cells will replace the active TP cells with individual cells that represent these combinations.
At all times at least 1 TP cell will be active.
The design has 2 key elements:
- Propagation of TP cell activity based on first-order modelling of historical transitions between SP cells.
- Assumes state splitting has created redundant SP cells representing the same FF input patterns, thereby allowing variable-order modelling using only first-order relations between SP cells.
- Inhibition of TP cell activity after a prediction failure.
Pseudocode
Variables:
nCells: The number of SP and TP cells
spCells: An array of SP cell activation values, length nCells
inCells: An array of TP cell inhibition values, length nCells
tpCells: An array of TP cell activation values, length nCells
spBest: The winning SP cell that matches the current FF input
spPrev: The previous winning SP cell.
spPred: The SP cell predicted to be the next best cell.
w( i, j ): Probability of transition from SP cell i to SP cell j in local graph
Note that weights w(i,j) represent conditional probabilities w(i,j) = P( S'=j | S=i ) where S is the currently active SP cell and S' is the next active SP cell. The weights can be approximated using historical observations of transition frequencies between active SP cells. Learning of sequence memory transitions is not covered in the pseudocode below. Similarly, the process for learning and defining the set of SP cells is not described.
// clear tp cell activity after bad prediction
if( spPred != spBest ) { // last prediction wrong
inCells[ spPrev ] = 1;
tpCells[ all ] = 0;
}
// clear tp cell activity when we reach inhibited cell
if( inCells( spBest ) ) {
tpCells[ all ] = 0;
}
// clear inhibition of winning cell and activate it
inCells[ spBest ] = 0
tpCells[ spBest ] = 1
// now propagate this activity forwards
sp1 = spBest;
int i = 0;
while( i < nCells ) { // or stop when stable
sp2Max = null;
wMax = 0;
// find the most likely transition starting at sp1
for( sp2 = 0; sp2 < nCells; ++sp2 ) {
if( sp1 == sp2 ) continue;
if( w( sp1, sp2 ) > wMax ) {
wMax = w( sp1, sp2 )
sp2Max = sp2;
}
}
// check inhibition, and consider replacing
// sp1 with sp2Max to continue propagation.
if( sp2Max != null ) {
if( inCell[ sp2Max ] > 0 ) {
break; // don't propagate to this cell, it is inhibited
}
else { // not inhibited
tpCells[ sp2Max ] = 1; // activate
sp1 = sp2Max; // now project forward from here
}
}
int i = 0;
while( i < nCells ) { // or stop when stable
sp2Max = null;
wMax = 0;
// find the most likely transition starting at sp1
for( sp2 = 0; sp2 < nCells; ++sp2 ) {
if( sp1 == sp2 ) continue;
if( w( sp1, sp2 ) > wMax ) {
wMax = w( sp1, sp2 )
sp2Max = sp2;
}
}
// check inhibition, and consider replacing
// sp1 with sp2Max to continue propagation.
if( sp2Max != null ) {
if( inCell[ sp2Max ] > 0 ) {
break; // don't propagate to this cell, it is inhibited
}
else { // not inhibited
tpCells[ sp2Max ] = 1; // activate
sp1 = sp2Max; // now project forward from here
}
}
++i;
}
So what does it do?
Let's work through a simple example that includes deterministic sequences and a non-deterministic fork. Obviously, the latter cannot always be predicted.
Figure 1 shows a test problem where each state has a corresponding colour observation. Four different colours are observed (red, green, blue and black). Arrows indicate transitions between states. The "true" graph of the problem is shown to the right. On the left is a first-order graph of observations. State-splitting is necessary to build a more useful model of the transitions between states, creating redundant copies of states with red, green and blue observations. Let's just assume that the SP cells have formed the "correct" representation of the world shown (figure 1, right) because that learning process is too complex to explain here. The fork at state 1 is random, with a slight bias making transition from 1 to 2 more likely than from 1 to 5. State 1 is shown twice for improved presentation.
Figure 2, above, shows the annotations and styling we use to present the state of SP and TP cells in the following walkthrough of the proposed temporal pooling algorithm.
Figure 1 shows a test problem where each state has a corresponding colour observation. Four different colours are observed (red, green, blue and black). Arrows indicate transitions between states. The "true" graph of the problem is shown to the right. On the left is a first-order graph of observations. State-splitting is necessary to build a more useful model of the transitions between states, creating redundant copies of states with red, green and blue observations. Let's just assume that the SP cells have formed the "correct" representation of the world shown (figure 1, right) because that learning process is too complex to explain here. The fork at state 1 is random, with a slight bias making transition from 1 to 2 more likely than from 1 to 5. State 1 is shown twice for improved presentation.
 |
| Figure 2: Key for diagrams used in the rest of this post. We have 2 sets of cells: TP cells (left column styles) and SP cells (right column styles). |
Figure 2, above, shows the annotations and styling we use to present the state of SP and TP cells in the following walkthrough of the proposed temporal pooling algorithm.
 |
| Figure 3: SP cell 1 is active. Each SP cell has an associated TP cell (none active yet). |
Figure 3: Say that via state-splitting, we have constructed a set of 7 SP cells (centre graph, coloured and numbered circles). This means we also need to create 7 TP cells (we specified earlier that there would be 1 TP cell per SP cell). Each TP cell is associated with one SP cell. The magenta outline of SP cell 1 indicates that this cell has been selected as best matching the current FF input. None of the TP cells are active. The heavier arrow from SP cell 1 to 2 represents the higher probability of this outcome.
 |
| Figure 4: TP cells active via prediction. |
Figure 4: According to the algorithm given above, we will first activate TP cell 1 due to activity of SP cell 1 (magenta). Then, we propagate forwards the activation through TP cells 2,3,4 because this is the most likely path (brown outline highlights). Activation propagates back to TP cell 1 and then has no further effect, since TP cell 1 is already active. TP cells 5,6,7 are not activated because this is not the most likely path from TP cell 1. As a result, of our 7 TP cells, 4 are active {1,2,3,4} of {1,2,3,4,5,6,7}.
 |
| Figure 5: Remember activation of TP cell 1. |
Figure 5: The external world moves from a state generating the black observation, to green. SP cell 2 is correspondingly made active. The temporal pooler cells are now updated, starting by activating TP cell 2 (although it is already active). TP cell 1 remains active for two reasons: First, activity is not cleared unless there is a prediction failure. Second, activity propagates from TP cell 2 through TP cells 3,4, and 1, back to 2. Again, TP cells 5,6,7 are not active. There has been no change in the set of active TP cells as they are following a predictable sequence. In fact, the set of active TP cells will not change while visiting any of the active states {1,2,3,4}.
 |
| Figure 6: Prediction failure. We predicted a transition from SP cell 1 to 2, but observed from 1 to 5. Inhibit activation of TP cells associated with SP cell 1. |
Figure 6: If, randomly, a transition from SP cell 1 to SP cell 5 occurs, the pattern of TP cell activation will change. TP cell 1 will be inhibited (magenta X) due to the prediction failure. TP cell 5 will be activated and the activity will propagate through TP cells 6 and 7. Crucially, it cannot propagate further due to inhibition of TP cell 1. This prevents the set of active cells extending to TP cells 2,3,4.
 |
| Figure 7: TP cell 5 activity remembered. TP cell 6 directly activated by SP cell 6. TP cell 7 activated by prediction. TP cell 1 inhibited. |
Figure 7: For the remainder of the sequence R-G-B, TP cells 5,6,7 remain active. While Green is observed, as shown above, TP cell 5 remains active not due to forward prediction but due to the fact that TP cell activity is retained until there is a prediction failure.
Discussion
In this example, TP output is one of two subsets of cells, either {1,2,3,4} or {5,6,7}. In the next hierarchy layer, the active subsets would be associated with specific SP cells that would represent these sequences of observations in their entirety.
This is a relatively simple scheme that only requires inhibition and prediction. A prediction failure is always followed by a change in TP output, due to the inhibition mechanism, even if the most likely path leads back to the original set of active cells prior to the failed prediction. Recall that the local graph being modelled in a region of cortex will always feature cycles and have no terminal states. Without inhibition, we would end up with scenarios where all TP cells were constantly active regardless of which fork was taken (if a lower-probability outcome occurs). This would obscure the unpredictable transitions and make them unavailable for modelling higher in the hierarchy.
At least 1 TP cell is always active (the cell that is associated with the currently active SP cell).
The output of the temporal pooler is maximally stable when sequences of observations are predictable. This is ideal if we assume the predicted outcome is the most common outcome. The propagation mechanism as proposed makes best use of predictable transitions - sequences of arbitrary length can be replaced with a constant output, if the system is sufficiently predictable.
Note that this method can only guarantee to simplify the observed problem in combination with state-splitting. The latter is required to build more and longer deterministic sequences of SP cell activation, even if these sequences are not predictable. It is assumed that given increasingly higher-order interpretation, predictability will be achieved.
In a future post we will look in more detail about the process of state-splitting.


No comments :
Post a Comment